Notice. Since this seems to be such a common question I think I should describe my background. I have a PhD in molecular microbiology and I was once a tenured academic (professor). I now work in the biotech sector. I have published a number of peer reviewed papers in virology. If you really want to know more about me then visit my About Me page.
Like nearly everyone on the planet I am worried about COVID-19. SARS-CoV-2 (the virus that causes COVID-19) appears to be killing between 1% to 3.5% of the people we know it infects (i.e the case fatality rate) and has a R0 (i.e. how many new people each person infected goes on to infect) of between 2.5 to 3.9 or even higher.
Newer reliable serology data, such as the studies done in Switzerland and China (i.e. not Santa Clara or Los Angeles), suggests the true infection fatality rate (IFR) is around 1%. Left to run wild, this virus will kill tens of millions of people worldwide.
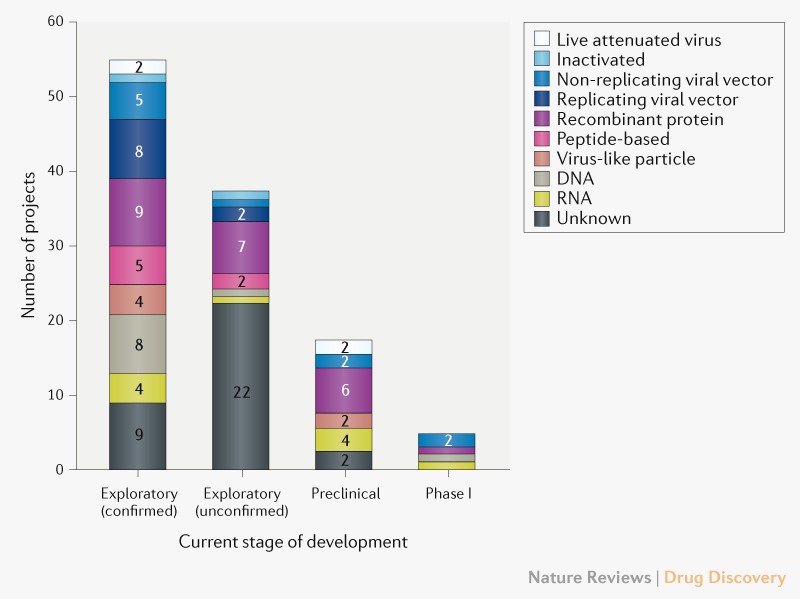
Many governments of the world have implemented strict population isolation protocols to try and limit the spread of the virus, but the economic cost of this is extremely high. A vaccine for COVID-19 is 12 to 18 months away (at best).
We are stuck in a diabolic situation where the only way to prevent the economy sliding into a slump deeper than the Great Depression is to consign many tens of millions of people to an early grave. Is there a way out?
SARS-Cov-2 Viral Diversity
SARS-CoV-2 like all viruses mutates (changes) overtime. Many of these genetic changes are small (single nucleotides) that are not important to the replications or transmission of the virus from person to person, but they can be used to identify the origin of the virus. DeCODE genetics for example has been testing Icelanders for COVID-19 and genome sequencing the SARS-CoV-2 strains isolated. They (and others) have found two very important pieces of information:
- More than 50% of the people infected with SARS-CoV-2 are asymptomatic or have only a mild case (i.e. they have no serious illness).
- They can identify the geographical origin of the strains by the genetic differences (mutations) between the different strains.
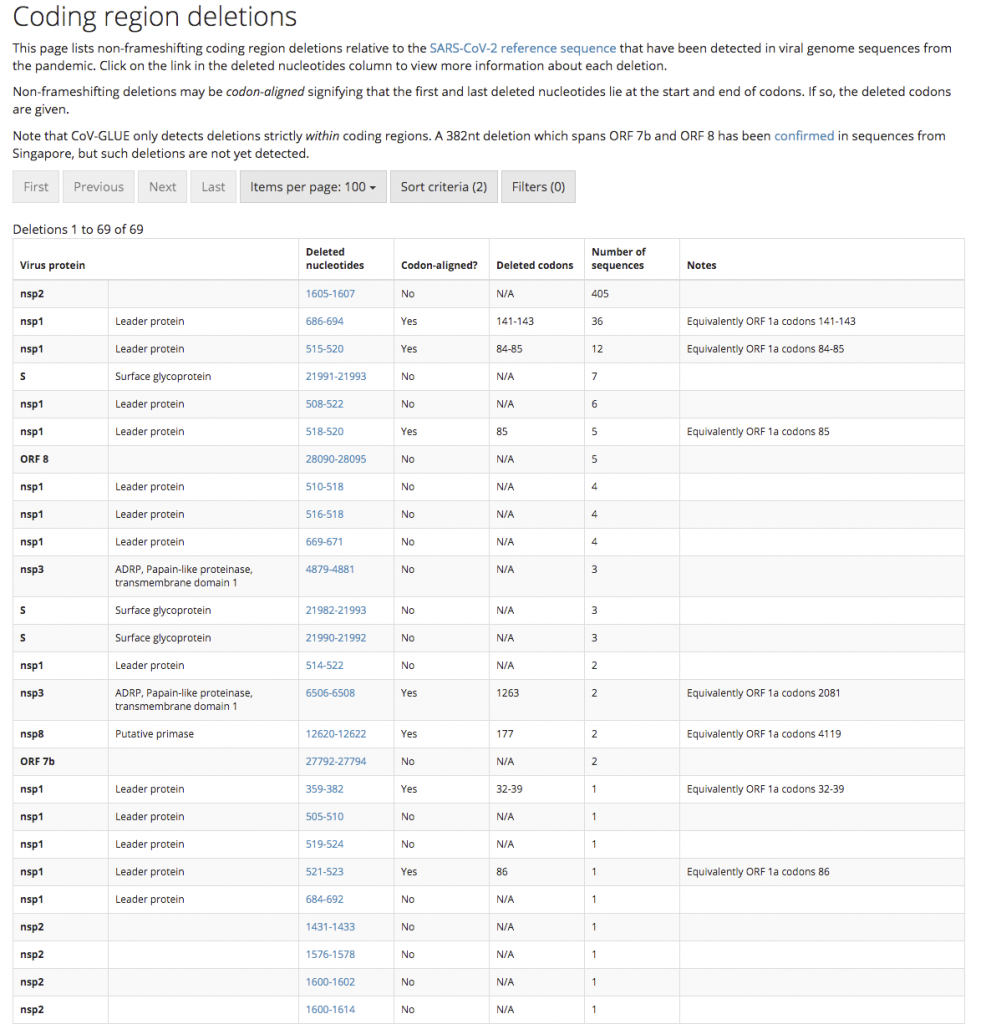
Furthermore, researchers in China have identified a mutant strain of SARS-CoV-2 which appears to be less pathogenic than most strains infecting people. The strain (ZJ01) had single nucleotide mutations in a key functional gene that made it less able to spread through the body. A recent pre-print paper from Singapore has described a mutant strain of SARS-CoV-2 with a 382 nucleotide deletion that was found in a cluster of patients in one of their hospitals. An even more recent paper from China found 11 of strains of SARS-CoV-2 with variation in pathogenicity in cell culture.
This data suggests a simple and testable hypothesis – there are natural strains of SARS-CoV-2 in the world that have mutated to be non-pathogenic (asymptomatic or mild), but which are still infective and will provide immunity to the more pathogenic (deadly) strains.
If we can find one of these non-pathogenic viral strains out in the wild we could give it to everyone in the world and solve our diabolic problem. This non-pathogenic (attenuated) strain would act much like the live attenuated (oral) polio vaccine.
Update. The genome sequence data collected to date has found that there are many SARS-CoV-2 strains with gene deletions.
How do we find an attenuated SARS-CoV-2 strain?
This hypothesis is worthless if we have no way of finding any of these non-pathogenic SARS-CoV-2 viral strains. Luckily there is a quick and cheap way to find these strains if they exist – test asymptomatic/mild case swab samples for COVID-19 and then genome sequence the SARS-CoV-2 strain that has infected them with the aim of identifying a virus with mutation(s) in essential viral gene(s). This is what the Chinese researchers did to find their less pathogenic strains and how the Singaporean researchers found their deletion mutant strain.
This approach is cheap (a couple of hundreds of dollars per virus strain) and quick (a week or less). With little cost we could sequence a few thousand viral strains, or even tens of thousands of strains, from positive swab sample from asymptomatic and/or mild case until we find a virus strain with the right mutations to make it harmless and which could work like a vaccine to protect us from the dangerous strains. We would know from epidemiology that this strain can still reproduce in people and lead to immunity, but not make people seriously ill.
Update. I have written a step-by-step post on one way we might go about putting this proposal into action.
What viral mutations are we looking for in a good non-pathogenic viral strain?
We would ideally be looking for a virus strain with a large(ish) deletion in an essential viral gene like the strain found in Singapore (unfortunately this strain is too dangerous to use as it put people into hospital). This sort of deletion mutation is easy to spot in the SARS-CoV-2 genome data, and because the genetic information has been removed, it makes the virus unlikely to be able to mutate back into a dangerous strain. Ideally, the strain identified will have infected a large number of other people in the local area so we can know it is safe from the epidemiological data. This will be important for getting regulatory approval to use the strain.
Has this ever been done before?
Yes. The polio, measles, rubella, mumps, and chickenpox vaccines are all live attenuated viruses. Even something as dangerous as smallpox was controlled in the 18th century using a variation of this idea called Variolation. The idea was the doctor would deliberately infect you with a less harmful strain of smallpox (often at a low dose) to make you immune to the more deadly strains of smallpox. Of course, they didn’t know how this approach worked in the 18th century, but it was still very effective and millions of people were saved from dying from smallpox by it.
Some people have been calling using a low dose of the virus Variolation, this is not what we think Variolation was, but this is a complex topic. While such a low-dose approach of the dangerous strain might make the COVID-19 less dangerous to the person being deliberately infected, it doesn’t make the virus any less dangerous for those around you that you might infect later. Such an approach could also not be used on the vulnerable, leaving them exposed to the illness. There is no reason in principle that the low dose idea couldn’t be combined with the attenuated strain idea and it might even be a very good idea.
In regards Corona viruses, there was considerable work done on determining which regions of the SARS-CoV can be deleted to create an attenuated virus that provided protection from the wild type dangerous version. Jose Regla-Nava and colleagues identified that SARS strains with gene deletions in the E protein were attenuated and provided good protection from later infection with SARS-CoV. Furthermore, these researchers found the deletion strains were genetically stable when grown in cell culture. SARS-CoV-2 contains the same E protein and deletions in the E protein gene may provide the same attenuation.
What are the risks?
The major risk is a mutant virus we think is safe is not 100% safe. While we can use community spread of the identified strain to estimate how safe it will be (i.e. if it has infected 1000 people and none have got seriously ill then we should have a pretty good idea that it is safe), our knowledge will be incomplete. We can of course spend the next few years testing and trialling, but if we do this by the time any strain is shown to be 99.99% safe (not even the polio vaccine is 100% safe) we will have all got COVID-19 and the world’s economy will be a smoking ruin.
We have a choice of taking some risks now, or face the certainty of a much worse problem later. Time to accept some risk and do something now.
Q & A
I have been getting a few questions on this post so I thought I would address them here.
How do you know there is an attenuated viral strain out there?
Because such strains have already been found. I am hypothesising that there is more than one based on the known mutation rate of coronaviruses and the number of cases. Coronaviruses like SARS-COV-2 mutate continuously (this is why companies like deCODE can tell the geographical origin of different strains) as the molecular machinery for replicating their RNA genome is not very accurate, although it is more accurate than the machinery of other RNA viruses like HIV. When you combine this with the millions of mild cases out in the world, the odds are on our side that there is at least one person infected with a strain that has a mutation that makes the virus less dangerous (attenuated). We just need to go and look for this strain – luckily the tools we need to use (genome sequencing) are now cheap and quick. What would have been impossible 20 years ago can now be done in a week.
Aren’t most people who have mild/asymptomatic cases infected with a dangerous strain?
Yes. Almost all people (>99.9%) who are infected (and have a mild case) are infected with a dangerous strain of the virus, they just happen to have an immune system that can control the virus well. With COVID-19 a mild case does not mean you are infected with an attenuated strain – for most people with a mild case if they happen to infect a person with preexisting conditions or who is old, that person will be at a high risk of dying. A mild case does not equal a harmless strain.
My argument is coming from the other direction. While almost all mild cases of COVID-19 are caused by a dangerous strain of SARS-CoV-2, an attenuated strain of SARS-CoV-2 will only cause mild disease. If you want to find an attenuated strain you need to look at mild cases even though >99.9% of the people you check will be infected with a dangerous strain. What we want to find is one of the rare natural viral mutants that has a mutation that makes it attenuated. Where you will find such a viral mutant is in people with the mild/asymptomatic form of the disease.
I am NOT arguing that people with mild cases are infected with a mild strain of the virus. If this is what you think I am saying please take the time to read carefully what I have written – yes it might sound like this is what I am saying from a five second scan, but this is not the case.
Why only sequence those with mild/asymptomatic cases of COVID-19 in the search stage?
If I had to choose one aspect that gets most misunderstood by people who read this idea, it would be the reason for sequencing only mild/asymptomatic cases. This choice is purely an efficiency issue. In an ideal world we would sequence the strains from every case in the world, look to see if we can find mutants with deletions, and then check what was the clinical outcome of those infected with each strain was. If we find that all clinical cases of a particular mutant strain are mild/asymptomatic, and no cases ended up in hospital, then we would have our candidate strain.
Unfortunately we live in a constrained world where it is not possible to collect and sequence the virus from every case of COVID-19, Given this, where should we look first? Since we are looking for a mutant strain that only causes mild/asymptomatic cases, we can exclude patients in the first pass who have serious symptoms. Any strain that causes serious illness won’t be a strain we want to use.
It is only once we find a good candidate attenuation strain that has the right sort of mutation (a deletion) that we then collect swab samples and sequence all cases in the local area. At this point we will need to look at both the serious and mild cases (include everyone in the local hospitals), to get the full clinical picture of the strain. This approach of initially screening just the mild cases is a simple way to make the search process for an attenuated strain more efficient and practicable.
Can’t we just wait for a COVID-19 vaccine?
No. Apart from the time it will take to develop, trial, and mass produce a vaccine (12-18 months), it is unlikely that any vaccine will be practicable. The reason why is immunity to respiratory viruses (like corona) doesn’t last long – 6 months to 2 years. We would have to keep vaccinating everyone in the world every year (or maybe every 6 months if we are unlucky). This just isn’t going to work in the real world (especially poor countries) and is one of the reasons we don’t have a vaccine for the coronavirus strains that cause the common cold. Unless we can drive the current dangerous SARS-CoV-2 strains to extinction we are going to have a problem with this disease indefinitely.
Update. A very interesting pre-print has been released on the natural immune response to SARS-CoV-2. Many people don’t develop much of an initial immune response (IgM antibodies) and the long term immune response (IgG antibodies) fades quickly after just two months (see figure below). While this is more of an issue for those people arguing that we should lift social isolation restrictions to allow the population to develop herd immunity (this would just kill a lot of people and any herd immunity would quickly disappear), it is also a concern for any vaccine approach that can’t drive the dangerous strain to extinction (i.e. all the other conventional vaccine proposals). 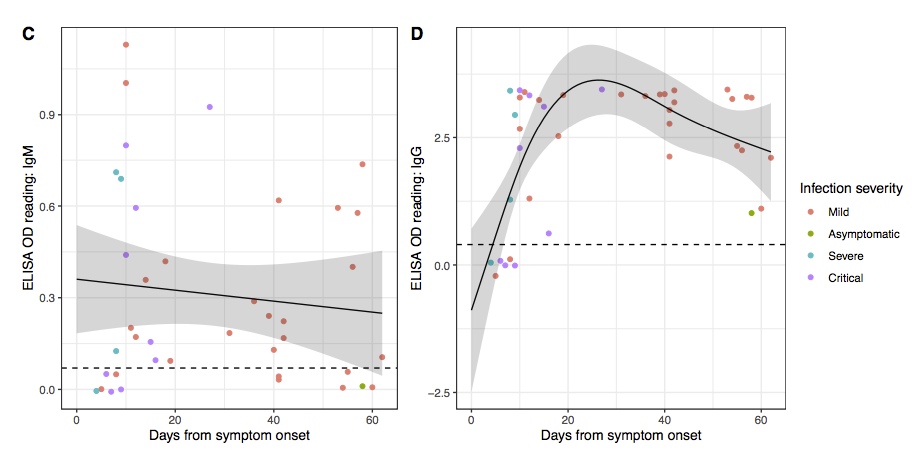
Won’t the mutations in SARS-CoV-2 make this proposal fail?
The coronavirus are relatively stable genetically for an RNA virus. The coronavirus strains that cause the common cold tend to not change much antigenically over time, unlike the Influenza, HIV and Hepatitis C viruses. The way the common cold strains spread is through our immunity to them declining fairly rapidly (in months, not years). The result is the same common cold strain can infect you multiple times in your life as your immunity to it fades, rather than it having to change antigenically so much that our immune systems doesn’t protect us anymore.
This is good news as it means we may only need to isolate one mutant strain. Even if in the worse case SARS-CoV-2 does change so much antigenically that our attenuated strain no longer offers protection, we can just repeat the process we used to find the first strain to find the next strain. More fundamentally, this is a problem for all vaccine approaches that we will need to deal with if it happens, but it shouldn’t stop us from acting now.
Update. A recent non-human primate study out of China using a conventional vaccine approach found that the antibodies produced were neutralising for all strains found around the world. It really does look like we only need only one attenuated strain to provide protection for all strains.
Isn’t social distancing and quarantining solving the problem?
Yes and no. Yes countries like South Korea and Australia have shown that through mass screening and social distancing you can keep a lid on the disease, but this leaves the population susceptible to a new outbreak. Singapore and Japan have recently seen this in action where they eased restrictions and found the disease came back and they had to reintroduce restrictions. I don’t think many people want to live for years with cycles of restrictions, easings and further outbreaks.
Wouldn’t the use of such an attenuated strain just be a vaccine?
Yes in one way, but it is a little more subtle. Assuming we can find an attenuated strain, then how to best use it a separate question. The most important thing to note is that a natural attenuated virus is not a vaccine. It is just a natural virus that you can catch in a natural way. Hang out with someone infected with the attenuated strain and you will catch it without doing anything, go home and those around you will catch it from you, and so on. While I wouldn’t suggest this is the best way to get an attenuated virus out into the community, such natural spread is outside the regulatory framework for vaccines.
Of course the use of a natural attenuated strain as a vaccine would fall under the regulations for vaccines, but the mere existence of an attenuated virus does not make it a vaccine. Each regulatory authority around the world would need to weigh the evidence of safety (obtained from the epidemiology) against the risks. Some regulatory agencies may decide the rewards from using such a strain as a vaccine is worth the risk, while others may decide they are not.
What effect would using an attenuated SARS-CoV-2 strain have on the dangerous strains?
Giving a natural attenuated viral strain deliberately to lots of people would change the ecosystem for the dangerous strains of the virus. The dangerous strains would find it difficult to spread through the community as many (most) people would have already been infected (and hence immune) with the attenuated strain (i.e. have herd immunity). Overtime the dangerous strains would become rarer, and the attenuated strains more common, until eventually the dangerous strains would become extinct and we would just be left with the mild version floating around. While we would not be able to get rid of this mild strain, it would just be another of the hundreds of viruses out there causing common colds. This proposal at its base is really one of replacing the dangerous strains with a less dangerous strain that we can live with.
It is the ecology aspect of this proposal that makes it different to other attenuated vaccine proposals. Currently all attenuated vaccines developed in the lab are designed to just protect the person receiving the vaccine. An attenuated strain identified by screening infected people (epidemiology) will identify a strain that can still be transmitted from individual to individual. This makes the use of such a strain radically different than a conventional lab created attenuated viral vaccine.
Why can’t we just use the less pathogenic SARS-CoV-2 strain already identified in China?
While the ZJ01 SARS-CoV-2 strain identified in China appears to be less pathogenic, the mutations that make it so are single base changes. These can easily mutate back to the more dangerous version of the virus. The viral strain we want to find will have a deletion mutation where a section of the viral genome is removed. Deletion mutations are much more difficult to mutate back to the dangerous type since rather than just a change from one nucleotide to another (e.g. C > T), the deleted region is missing and can’t easily be recreated by mutation. Put simply, deletion mutations are more stable to back reversion.
How should any natural attenuated SARS-CoV-2 strain be used?
This is not a decision for me. I think the regulatory authorities like the FDA will approve the use of a natural attenuated strain on the basis of the epidemiological data collected finding the strain that show it is safe. While most regulatory agencies are extremely conservative and slow, they are not idiots and they understand that using an approach with some serious unknowns might be better than the alternative of waiting for the perfect vaccine. COVID-19 is a problem with only hard choices.
I do think it would be impossible to prevent the use of any such strain even without regulatory approval. Assuming a natural “safe” strain is found and a test for it developed (this is relatively easy since you are looking for the presence of a deletion), people will start spreading it between themselves at a “grass roots” level. I think this outcome will influence the decision of the regulatory authorities to approve its use – better to have the attenuated virus spread in some controlled and regulated way, rather than illicitly by people doing it on their own. Just to make clear, I am not suggesting that it is a good idea to allow “grass roots” spread (I think it is a bad idea), nor am I advocating for it, just that I think it won’t be possible to stop such spread if the regulatory authorities don’t license it.
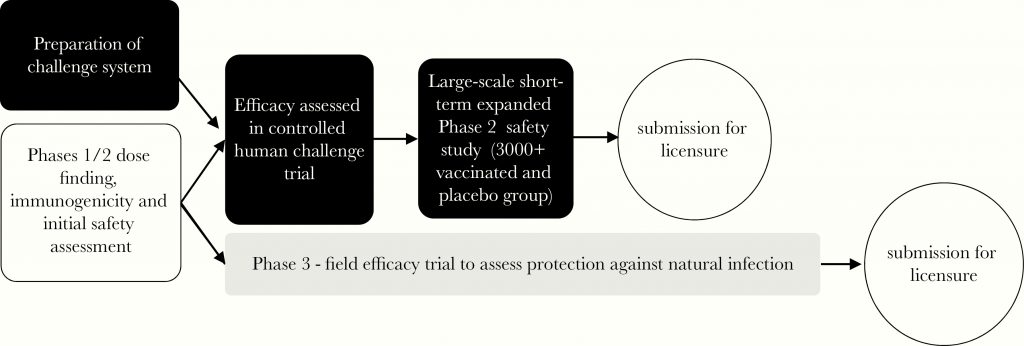
Even if you think a natural attenuated strain is too dangerous to use as a vaccine directly, it would be extremely useful to speed up other vaccine approaches. One way it could be used is to give it as a challenge to volunteers who have received another vaccine to see if the vaccine provides protection without putting them at risk. Knowing if a vaccine works (i.e. provides protection from the disease) is one of slowest steps in new vaccine development. If you are going to do a viral challenge with SARS-CoV-2, you really want to use a strain you know is harmless, rather than one you know could kill your volunteers. Of course to do this you first have to make the effort to find such a harmless strain.
I have objection X which means this whole idea is worthless!
If you are not a scientist then I will be blunt and say it is almost certain that your objection is either wrong or irrelevant (sorry for being harsh). While I have presented the idea in as non-technical way as is possible, it is still a complex scientific idea with many complex parts. What might seem to you a very strong argument is likely to not hold up when explored in depth.
If you are a scientist try not to get caught up too much in me skipping over certain technical details and instead focus on the bigger picture. This is a proposal for an approach that may work, not a grant application where all the experiments have already been done. There is a non-trivial chance that this approach may not work. I am aware of this, so I ask you to please focus not on all the possible ways it may not work, but if there is anything that will make it certain to not work. If there is some fatal flaw then please let me know, not that step x might not be easy, or we don’t know some particular fact. This is a problem where assumptions have to be made and risks taken.
It is not ethical to use such a natural attenuated strain!
This is a proposal to search for a natural attenuated viral strain of SARS-CoV-2, not an proposal to use such a strain. Finding such a strain does not mean we have to use it, just that we can use it. Ultimately the decision to use such a strain will be a political and social question weighing up up all risks, but we can only have this debate if we make the effort to find such a strain and show it is safe.
If you think that the whole idea is unethical under all circumstances, it is worth keeping in mind that it exactly the same as what Sabin did with the oral polio vaccine. He found a natural attenuated polio strain in a child with a mild case of the disease and used it in his vaccine. His vaccine has gone on to prevent tens of millions of cases of polio. Yes there was and still are risk with his vaccine, but most people think it was huge step forward for humanity.
How would this search work in practice?
I have written a second post describing how we might go about turning this idea into reality – How would a search for a natural attenuated SARS-CoV-2 strain work in practice?
Update. I have been trying to get this idea in front of someone with the clout to make this happen, but without much luck because the people with the clout (say Bill Gates) don’t read messages from random people like me. For this idea to succeed it needs a two hop process (i.e. someone knows someone, who knows someone, who Bill Gates will take seriously). If you think you might be that first hop person then please get in contact with me at daniel.tillett@gmail.com.
I should add that it doesn’t have to be Bill Gates that can push this forward. If you think you know someone who could help then get in contact with me. Unlike Bill I will respond to your email :)
Update 2. I have modified this post to better explain those aspects that have confused people (my fault). I hope this idea is now clearer. Keep the comments coming and if you think someone you know would be interested then pass the post along along.
Update 3. This is directed at my fellow scientists. If you happen to working on this exact idea please get in contact me – not least so I can give you lots of money to speed up your work.
Update 4. This field is moving at an amazing pace with new results coming out every day. I have updated this post with the most relevant new data.